Yesterday I introduced the main character to this drama of the hunt for the elusive bird leptin. Now its time to introduce the other players and get on with recounting how the discovery of the bird leptin gene was made. Several years ago I began collaborating with Dr. Rich Londraville (UA Biology) on a project involving the elucidation of the function of leptin in maintaining massive blubber layers in arctic adapted bowhead whales. Dr. Londraville has studied leptin for much of his career with a particular interest in fish. I brought a PhD candidate, Hope Ball (now Dr. Ball) into my lab to take on the bulk of the project (that is a story for another time). In joint lab meetings with the Londraville’s lab we often found ourselves talking about the diversity of the leptin hormone and its function across all vertebrates. If you don’t know the story of leptin then you should probably read Part I (The Elusive Bird Leptin: And Now for the Rest of the Story…) before continuing.
We live in a human and mammal-centric world with respect to our initial understanding of the function of many animal genes. Often this means we learn how something works in a human or mouse model and then hope that we can export our knowledge about how genes work to other animal systems. However, many times genes can take on different functions in other animals even those that may be closely related.
Leptin is a good example of a gene that has different effects across diverse animal systems. In Part I I said that leptin is expressed in fat cells of mammals. However in fish it is the liver that is the primary producer of leptin. This fact alone strongly indicates that fish likely use leptin in different ways than do mammals. So what are all the effect of leptin and which effects are likely to be of primary importance and which are lineage specific? Comparative biological studies versus human-centric or model organism studies can best provide the answers to those questions.
As comparative biologists our discussions of how leptin might be used in a variety of different animals and how it can be used to explain something like the biology of super-fat whales frequently veered off into asking the question“what’s up with bird leptin?”
Concocting a new approach to finding bird leptin
In the spring of 2013 UA Integrated Biosciences PhD candidate Jeremy Prokop and several other UA graduate students along with two undergraduate students in my lab who had been working on dynamic molecular simulations (ie. modeling) of leptins developed a new approach to searching for bird leptin. Armed with the idea (possible explanation #2 from Part I) that a second gene had taken the place of leptin as the activator of the leptin receptor allowing the ancestral leptin to be lost from the genome, they proposed to use our knowledge of the binding site of the receptor derived from our studies of leptins from many different animals to predict the basic protein shapes that must be made by this potential replacement gene. With that search image they would then model genes of unknown function or of possible duel-function genes from bird genomes in hopes of identifying a gene that could be producing a protein/hormone that could be doing the job of leptin in those birds.
An analogy to this replacement gene idea would be looking for puzzle pieces. The piece that has the “blanks” or “pockets” require a matching “tab” to fit it. A hormone (the tab) and a receptor (the blank) work in a similar fashion. We know the blank (receptor) exists in birds but can’t find the tab to fit it. But maybe the piece with the blank is all red and we are looking for a piece with a tab that is all red but that piece doesn’t exist but rather another piece with just a smidgen of red around the tab exists but most of the piece is actually yellow. What our students proposed is that for birds the original puzzle piece (leptin) is gone and has been replaced by another piece (gene) that looks nothing like the original piece except that the part that is crucial for linking to the blank has the same shape and color to make the connection. This would allow another gene/piece to take over the function of the original piece and connect to it.
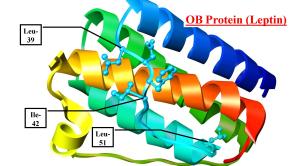
While a relatively simple-sounding idea it nevertheless was computationally very challenging. Only very recent advances in molecular modeling have allowed such examination of proteins and even so the time required for the simulations is significant (on the order of a day to several days per gene on a fast computer running at 100% cpu capacity!). This group of students wrote a research proposal and submitted it with myself and Rich Londraville as faculty sponsors to the University of Akron for consideration for student research funds. The proposal was funding providing several thousand dollars for the project.
Putting it into action: summer 2103
With a research plan, money in their pockets and dreams of finding the elusive bird leptin the project participants began implementing their ideas as soon as spring exams were done. But science has a funny way of not working on a schedule or according to anyone’s plans and this project would be no exception.
Shark leptin a key to finding bird leptin – The strange route to the discovery of bird leptin began in Mexico! Rich Londraville sent me an email from there while attending a meeting of the North American Society of Endocrinology at Juriquilla, Queretaro, Mexico. At the time he was listening to a talk by a scientist who was presenting results from their efforts to sequence the genome of the elephant shark. During the talk a link was shown where the raw sequences of the shark genome could be located and searched. Londraville sent me the email below by email while still listing to the remainder of the talk.
Sent: Wednesday, May 22, 2013 1:36 PM
To: Duff,Robert Joel
Subject: Greetings from Mexico
http://esharkgenome.imcb.a-star.edu.sg/
Leptin in rat fish?
Sent from my iPad
I was in my office at the time and was excited to get the chance to find a more ancient version of leptin. I also knew that it could help us solve some questions about some weird thing we see in fish leptin that no one has been able to explain. I immediately found the coelacanth (a lobe-finned fish different that most of your bony fish) leptin sequence that I had just retrieved from a genomic database a few weeks earlier. I used that sequence to search the shark database and got several weak matches. The best one had a 28% similarity to the coelacanth query sequence. Having looked at a lot of leptin sequences from many diverse organisms (the advantage of doing comparative biology again!), I recognized right away that despite such low similarity some important sequence elements of leptin where there.
I proceeded to take that best hit from the shark and use it to do a search on the billions of sequences found in the international sequence databases. That search resulted in closest matches to a frog and some land animal leptin sequences. The chance that this shark sequence would match up to a frog and animal leptin sequence is very slim and so I was pretty sure that we had found the first leptin gene in a shark. In fact this was the first definitive leptin sequence found in any organism “lower” than any bony vertebrate. I wrote back within an hour of receiving the original email:
Rich,
I took the hit I got from the Ceolacanth leptin and then blasted that sequence in Genbank and it came back with all hits to leptin with Xenopus being the most similar 28% (sequence identity but 49% amino acid property conservation) and then lizard at 26% with 99% query coverage. Can’t believe it, shouldn’t be that easy to find, right. BTW, NO fish in the top 100 hits (remember that Ceolacanth isnt’ in NCBI yet). This could show that fish lineages have gone wacky with leptin while its actually older and the lineage that led to land mammals retains the more conservative side of leptin.
I can see conservation of some of the same important amino acids in the alignments that NCBI is showing me (all the Cysteine residues that are conserved in all leptins are here in this elephant shark). Very cool!
I will send to Jeremy.
Joel
That evening I took the shark sequence and subjected it to molecular dynamic simulations and the next day I had confirmation that this sequence indeed produced a protein that folds into a structure that is easily recognizable as the leptin hormone.
Like stepping stones across a stream, one sequence can lead to the discovery of other similar sequences. With one shark sequence now in hand I was able to search another shark genomic database and find a “skate” leptin sequence. These new sequences could now be aligned with fish, amphibians and mammals to give us a new picture of just what parts of the leptin gene are conserved across the evolutionary landscape and what parts have been free to vary over time. Genes of greater phylogenetic (evolutionary) distance are very helpful in this cause and with the shark sequence now in my alignment I had some new ideas about what sequences could be used to search other genomes for leptin.
Alas, a quick search of the chicken genome yielded no significant similarities. I began to think that to find the bird sequence we would need to have more sequences from the animals that are most similar to birds. Unfortunately dinosaurs are extinct! The next most similar animals to birds are living reptiles and specifically members of the Archosauromorpha; reptiles which includes crocodiles, birds and dinosaurs. I soon discovered that there was a crocodile genome project underway and the raw sequence files were available.
Working with a billion bits of DNA sequence is not exactly in my comfort zone. Fortunately, two undergraduates in my lab, Cameron Schmidt and Donald Gasper, were working on the bird leptin problem from the original angle of protein modeling. Both were very adept at implementing bioinformatic methodologies. I tasked Cameron with downloading the genome files and learning how to set up a local search program so we could look for crocodile leptin. At the same time we also obtained sequences of the python (snake) genome and did the same search.
On June 5th I sent Cameron the following email after looking over all of my leptin protein sequence alignments:
Cameron,
Here is the most conserved section for Coelacanth/amphibian/reptile that I can find: QFLLPXNLKVSGLDFIPA 75% match to this would be worth looking at.
Joel
In retrospect I should have said a 40% match would be worth checking. That is a very short piece of protein sequence but I figured if he could locate that among the billions of pieces of code from these genomes then we would have found the core of the leptin gene. By the next day Cameron had found sequences in both the alligator and python genomes with a greater than 40% similarity. Some additional work confirmed that these were indeed leptin sequences and we were able to work out from that conserved sequence to get the entire gene.
I expect you can now predict our next step. With the python, alligator and a few additional reptile and amphibian sequences added to my fish, shark and mammal sequences I was able to make a new protein sequence alignment. While each individual sequence was quite different a pattern of conservative sequence motifs began to emerge. The one reptile and amphibian sequence that were originally available were wildly different from one another and therefore it was impossible to predict what sequences the bird genomes might have in common with them. But with these new sequences I sat down predicted what the bird sequence should look like with the assumption that it should be most similar to reptiles.
I sent a simple email to Cameron:
Search with this sequence NLKVSGLDFIPGERPXESLXXMDETLQVFQRILXSLPXEAXXXQIXXDXENLRSLLXLLGAXXGCXXXXXXXXXXXXNLTELLXXXPYTXXXVALDRLQKSLHXIXKHLDXLXXC
The X’s in the sequence represent positions where there is so much variability among leptins in my alignment that there was no way of telling what a bird might have. But the other letters represent a specific set of amino acids in linear arrangement that I thought were likely to be present in a bird leptin if they had one at all.
By now Cameron and Donald had downloaded and prepared two very recently available genomes of the Saker and Peregrine falcons plus there was the older chicken genome and an available genome of the zebra finch.

I walked down to the lab to see how the search was proceeding. Cameron Schmidt compared the short amino acid sequence that I sent him to the over 1 billion bases of code in short DNA sequence fragments of the Peregrine falcon genome. The search yielded numerous partial matches that were anywhere from 15 to 30% similar but the top match had a specific set of amino acids that looked very familiar to me. I literally jumped off my seat and raised my hands above my head like a field goal kicker after he watches the football sail through the uprights and said “we’ve found it!!” I ran around the lab I was so excited. When I finally calmed down I asked Cameron to do the reverse search with the short sequence from the falcon against the NCBI (National Center for Biotechnology) database. What we got back were hits to the turtles and mammals that were about 40% similar but clearly identified as leptin sequences. At this point I was pretty sure we had found leptin in a bird.
I quickly threw the sequence we found of the falcon in with my other sequences and produced a quick comparison to visualize the differences and similarities between them. Below is one of those first genetic similarity trees that I shared with my colleagues.
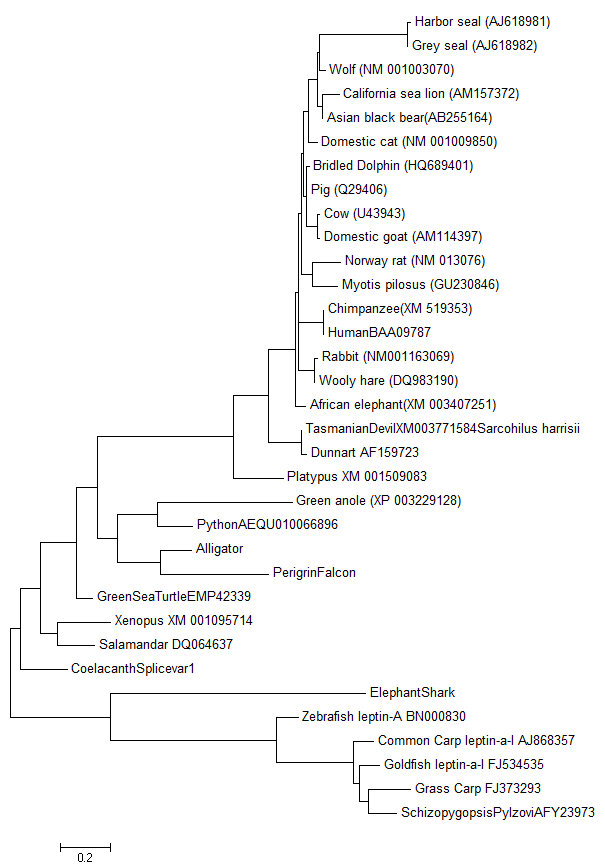
That evening (June 7th) I sent the following email to Jeremy Prokop:
Jeremy,
A very exciting day here. We think we have found bird leptin in the Falcon and Zebra Finch genomes. We will get together next week to talk about what to do to get this published quickly. Still lots of questions about why it isn’t in Chicken etc… Joel
Now that we had a bird sequence we could use that bird sequence to search for other similar sequences in other bird genomes. The first was the Saker falcon genome. We quickly found a similar sequence to the Peregrine falcon and then in zebra finch. A few days later we were also able to find pieces of leptin sequence from genome projects for the ground tit and duck. Each of these sequences was quite different from one another but more similar to one another than to any other leptin sequence from other animals. That pattern of divergence from non-birds yet similarity among birds is exactly what we hoped and expected to see from a putative set of bird leptins.
With some sequences in hand I could do alignments of them with other animals and see how different they really were. At the same time, Jeremy Prokop, now a PhD research associate at U of Wisconsin, Madison, who was spearheading the overall project took the sequences and began to do intensive computer modeling of the leptin hormones from the predicted protein sequences to test whether these leptin hormones looked like functional leptins and could bind to the receptor proteins that were already known.
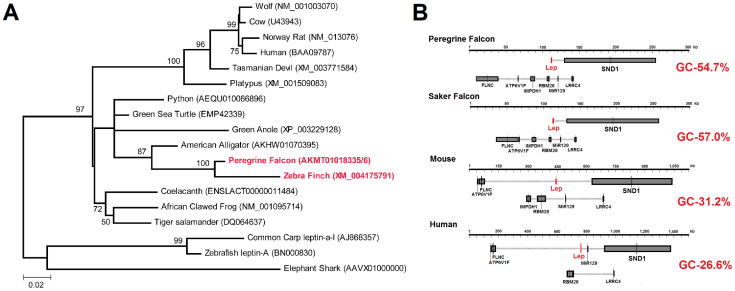
The rest of the story is in the paper published by PLOS ONE the last week of March, 2014. That manuscript if full of data analyses performed after we identified the bird leptin gene most of which were performed by first author Jeremy Prokop along with the other students. Additional collaborators also got involved to test other aspects of this bird leptin. Altogether the work resulted in the following basic observations:
1) Leptins from birds are clearly more similar to each other than to other animals – as expected. As a group they are most similar to alligators as predicted by evolutionary studies of birds.
2) Bird leptin sequences have much higher GC content (more G and C parts of DNA) than mammal leptin sequences. This is part of the reason that their overall sequence similarity is so low and their structures could be composed of such different amino acids while still retaining the same basic shape as mammals and other vertebrates.
3) Sequences on either side of the falcon sequence revealed that the SAME genes are found in the vicinity of leptin as are found in mammals and most other vertebrates. This means that leptin is found in the same part of the genome as it is in mammals.
There is much more data and work that went into the paper. If you are interested check out the original paper here.
Where do we go from here?
What will the future bring? I don’t know but there isn’t any going back, just new discoveries lying before us. What I do know is that there are plenty of mysteries to be solved. Stay tuned for more discoveries.
—————————————————————————————————————————–
A few addenda to my bird leptin story:
1) Chicken leptin: It is still a mystery! Where is the chicken leptin? We already knew that the position where leptin should be in the chicken genome is absent from the genome reconstructions so it was not surprising that our search using bird leptin yielded nothing from chickens. Does that mean that chicken has no leptin? Not necessarily.
I said earlier that the complete chicken genome doesn’t have leptin and the portion of the genome where leptin would normally be is absent in chicken. But is it really absent? I don’t think so. Just because the genome has been sequenced and called complete doesn’t mean that we really have the ENTIRE genome sequence. Most people don’t realize that no animal or plant nuclear genome has been 100% sequenced. The human genome is considered finished but it is really only 99 to 99.9% complete. There are portions of any genome that are very difficult to sequence either because they contain repetitive DNA that is hard to interpret or the chromosome is highly resistant to the cloning process used to prepare DNA for sequencing.
The fact that it isn’t just leptin that is missing from chickens but 10 or more genes that are in the same region as leptin in other birds and mammals makes me strongly suspect that this missing sequence isn’t missing because it doesn’t exist but rather it is missing because of difficulties in sequencing this region of DNA. I expect that more advanced DNA sequencing techniques that are in development will eventually sequence the genome and “find” the missing DNA sequences.
And yet the chicken genome is very perplexing because it isn’t just the DNA that is missing but all signs that the hormone is produced are not present including searches of EST (expressed sequence tag) databases also yielded no chicken leptin. The latter is a database of sequences that represent genes that are expressed in chicken tissues. Even if we can’t sequence the part of the genome where the chicken gene is one would think that we would see the chicken sequence being expressed as RNAs if it is being used to make leptin hormone.
So leptin has been found in birds but the chick and turkey leptin still remains elusive and an open question as to its existence.
2) Students make it happen! I’ve been talking a lot about what I did in this search for leptin but I was only a part of the team. You may notice that I am the fourth author on the paper. The bulk of the manuscript was written by a former PhD candidate Jeremy Prokop. Jeremy and other graduate students and undergraduates played large roles in both collecting and interpreting the data. This leads me to one final point:
3) Science does not always follow predictable lines of progress. Sometimes you have to be in the right place at the right time. Honestly, this is one of the discoveries that someone was going to make in the not too distant future. Leptin was not bound to remain elusive for long. With more and more genomic sequence information it is obvious that any number of people would eventually have related a sequence in one of these new bird genomes to leptin at some point in the near future.
This project started with a research proposal that outlined a specific strategy for finding bird leptin. A LOT of work went into the implementing that strategy but in the end, that email from Rich from Mexico set off a line of inquiry that eventually led us to find the bird leptin in a way that was not anticipated. Had the other research project not been underway I don’t think that we would have found the leptin because it took the expertise and experience of that project to allow us to the searches of other genomes and find leptin there and to be able to search the falcon genome. In other words, if it were just me I would not have been able to make much progress at all even if I had the right ideas.
Addenda:
NCBI: BLAST page http://blast.ncbi.nlm.nih.gov/Blast.cgi
Ensemble Genome Viewer: http://useast.ensembl.org/index.html
DNA and AA sequences:
Ceolacanth fish Leptin DNA sequence: >gi|557014814|ref|XM_006007836.1| PREDICTED: Latimeria chalumnae leptin-like (LOC102366460), mRNA ATGAATTCTCTGCTGTTGCATGTCTTTGGCTTACTGTGGATATTGATACCACTGTGCTCCAGCCGACCTG CCAAGATTGAAAAAGTCAAGAGTGATGCGAGAAATCTTACTCGAATTATAATAACCAGAATCCAGCAACA CCCGAACCAGTTCTTACTCCCTCTCAACTTGAAAGTATCTGGTTTGGAGTTCATACCTGCAGAGAGACCA CTGGAGAGCTTGGGATCCATGGACGAGACATTGGAGATCTTCCATTGGATACTGTCTAGTCTTCCCGTAG ACGACGTCACCCAGATCCTCTGGGACATAGAGAATCTGCGGGCCCTTCTCCAAACTCTTGCCACCACCAT GGGCTGTGAGCTTCACCAGACCACTGAGCTGGAAACCTTAAAGGACTTGGCCAAGGAACACTCCACGTCA CCTTACACCACGGAGAAGGTTGCCCTCGACAGGCTTCAGAAGTGCCTTCTCACAATGGTTAAGGAGCTTG AGCAGATCAAAGACTGTTAA
Ceolacanth fish Leptin Protein sequence:
>gi|557014815|ref|XP_006007898.1| PREDICTED: leptin-like [Latimeria chalumnae]
MNSLLLHVFGLLWILIPLCSSRPAKIEKVKSDARNLTRIIITRIQQHPNQFLLPLNLKVSGLEFIPAERP
LESLGSMDETLEIFHWILSSLPVDDVTQILWDIENLRALLQTLATTMGCELHQTTELETLKDLAKEHSTS
PYTTEKVALDRLQKCLLTMVKELEQIKDC
Conserved Leptin core protein sequence for tetrapods:
NLKVSGLDFIPGERPXESLXXMDETLQVFQRILXSLPXEAXXXQIXXDXENLRSLLXLLGAXXGCXXXXX
XXXXXXXNLTELLXXXPYTXXXVALDRLQKSLHXIXKHLDXLXXC
Elephant Shark Genome BLAST page: http://blast.imcb.a-star.edu.sg/cgi-bin/blast/blast_shark.cgi
Elephant Shark Leptin Protein Sequence:
S-Q-FKPLPSMRISG—–LERIPNTVSLS-GPDSLGPTIEGLHCYVEVL–RS-LGLENTAQIQADVEGLWLVLKSVAQS–WGCSEPQCSAAQQLITFLRNNTSYKL-TISHVVLEKLISYLQQAVRVLDNA-SRC
Perigrin Falcon DNA protein sequence:
MRWP-GMSLWGVLWLWLRPARGA-P–VRLEKVRADTRNLTRTLSARIQQLQA-R-QLFPLSLKISG—–LEAIPGEGAPE-GLGAMDHRLQLFQRLLGGL—A-AGGLPLAQIANDMENLRSLLAALAAP–PGCPLPRAPPAP-PG-LPDLLAEAPH-TAAGLALARLRICLDGIAARLDSL-PAC
Your similarity tree in this blog post is pretty much as expected, but the one in fig. 1A in the PLOS One paper seems weird in that it finds coelocanth leptin (representative sarcopterygian?) closest to amphibiia and closer to diapsida than synapsida, where you might expect it to be “basal” to tetrapods in general, as in the earlier tree. Is this just a trick of the light or how does it work?
LikeLike
Excellent question. There are a number of factors at work in the construction of the trees. The first tree was a very quick distance analysis (neighbor-joining) which is a fairly straightforward estimate of genetic distance but doesn’t account for GC/AT content, transitions vs transversions or make any assumption about evolutionary mechanisms. The tree from the paper was a maximum likelihood analysis and we did some work to determine what model of evolution would be best for our data set. Despite all that, the main reason the two trees are different is the different taxa used in construction them. Because these samples are so different from one another the addition or subtraction of a sample can tweek the the entire tree. In the supplemental files there is a tree with more samples and some of the relationships are different. That coelacanth position doesn’t have a lot of support where it is and so it can bounce around depending on what other samples might have a few amino acids similar or different. The bottom line is that the ceolacanth really is very similar to amphibians and more similar to them than to other fish. I wish we had a lungfish leptin but that genome is FAR larger than the ceolacanth genome and so they started with the ceolacanth before moving on to lungfish which I am sure will be eventually.
LikeLike
Oh yeah, I should have added that the lack of many fish in the outgroup really effects the placement of ceolacanth in the analysis. We wanted to keep the tree small so the birds wouldn’t get lost and because the original formatting was for Science and couldn’t take much space.
LikeLike
Thanks for that detailed explanation. Lungfish would presumably help to anchor coelocanths… somewhere. Is there any chance that extant tetrapods are polyphyletic, with modern amphibia closer to surviving sarcopterigia than amniota?
LikeLike
Yes, lungfish will probably help. There is still a chance they are polyphyletic but the majority of the data wouldn’t support that presently.
LikeLike
“So leptin has not been found in birds but the chick and turkey leptin still remains elusive and an open question as to its existence.” Need to get that “not” out of there.
Will you express a bird leptin construct in E. coli or yeast to look at the protein, or is that too 20th century?
LikeLike
Thanks for the grammatical catch there – its fixed. Much to do in terms of benchtop work including expression of the protein in E. coli. Some progress has been made but the reason that it wasn’t in the paper is that there is still something funny going on with birds that makes working with this part of the genome very difficult. If it were like other leptin in other animals we would have had that data included in the paper which maybe would have made it a Nature or Science paper but we felt that publishing what we had was important and so the rest of the story will have to come out over time.
LikeLike